Co-eluting chromatographic peaks remain a frequent challenge for nearly every GC/MS analysis. Conventional deconvolution typically examines the XICs (extracted ion chromatograms) and often fails to determine the correct number of co-eluting compounds (this method will typically over-fit, i.e., return more spectra of “pure” compounds than is actually in the peak) which tends to produce “false” spectra and many false library matches. GC/ID tries to improve upon this approach by first using Principal Component Analysis (PCA) to estimate the number of components more confidently in a peak, and then attempt to deconvolve them into pure component spectra (minimizing overfitting and false matches).
However, when there is not enough retention time separation (difference) between co-eluting compounds, all conventional deconvolution approaches will fail. It is a mathematical certainty!
This is where SMD comes into play. SMD is a new approach to deconvolution which directly fits the mixture spectra with all possible library spectra using multiple linear regression to find which “pure” spectra provide the “best” fit to the mixture spectrum. Of course, there are two problems to solve, 1) how to identify potential “mixture” peaks and 2) how to select a reasonable subset of compounds from the library to perform the fit (otherwise finding the best fit using all compounds from the library becomes an intractable problem). To identify likely mixture peaks, we can use reverse search (see “The Importance of Forward and Reverse Search”). Once we do this, we can select a subset of the library to perform the fitting by finding compounds that have a high reverse search score, and that have a RI value close to the peak. While fitting all possible combinations of the selected spectra is still computationally intensive, modern computers can perform such a task rapidly by using modern, optimized programming techniques. Once the best 2, or 3 (in practice 3 compounds is a practical limit) spectra are found that provide the best fit, we return these spectra as the most likely candidates. So, unlike conventional search, you do not get a ranked list of the closest matches which can be viewed and overlaid.
However, to perform SMD efficiently, a fast, unfiltered search must be done, which is accomplished using Cerno’s CPS Search.
One of the many powerful features of GC/ID.
Find out more from the GC/ID brochure.
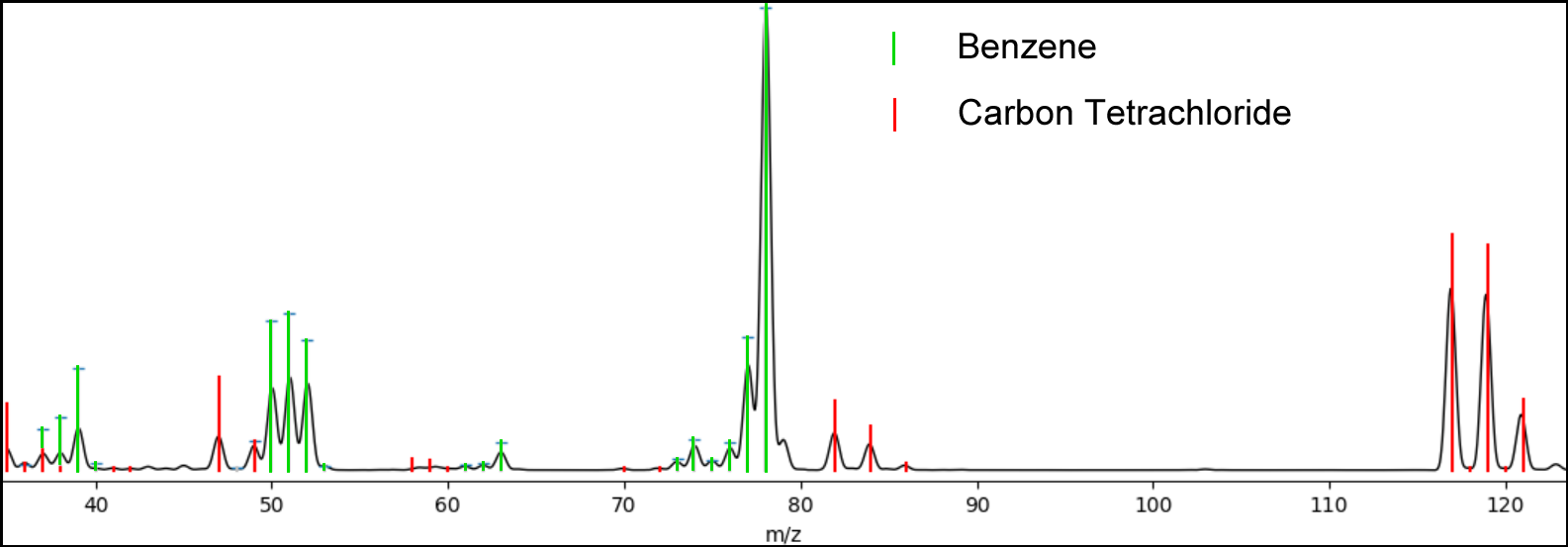
Figure 1. An SMD plot showing the search results from the coelution of Benzene and Carbon Tetrachloride. The black profile mode spectrum is a mixture of the 2 compounds, the green centroids are the library spectrum of Benzene, while the red represents Carbon Tetrachloride, and together they explain the mixture. Applying conventional deconvolution approaches, from any vendor, will fail in this case where the separation of the peaks is minimal (if any).